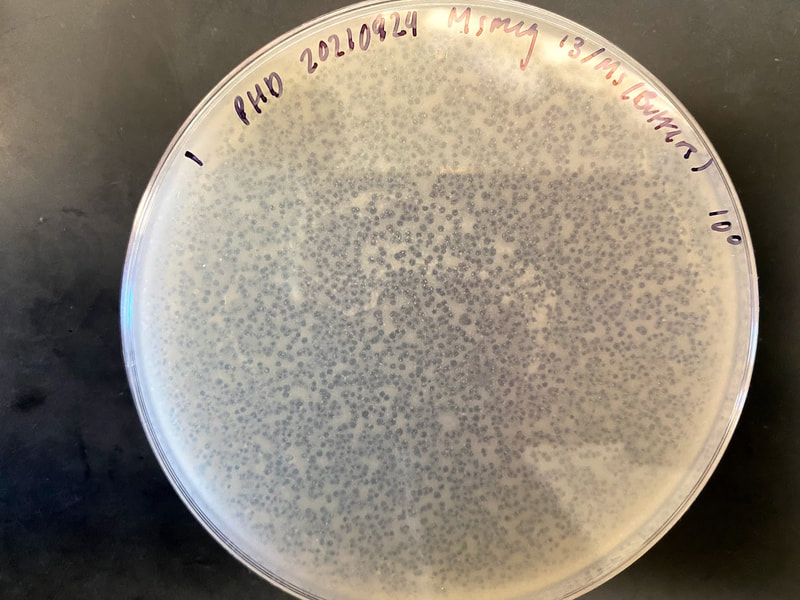
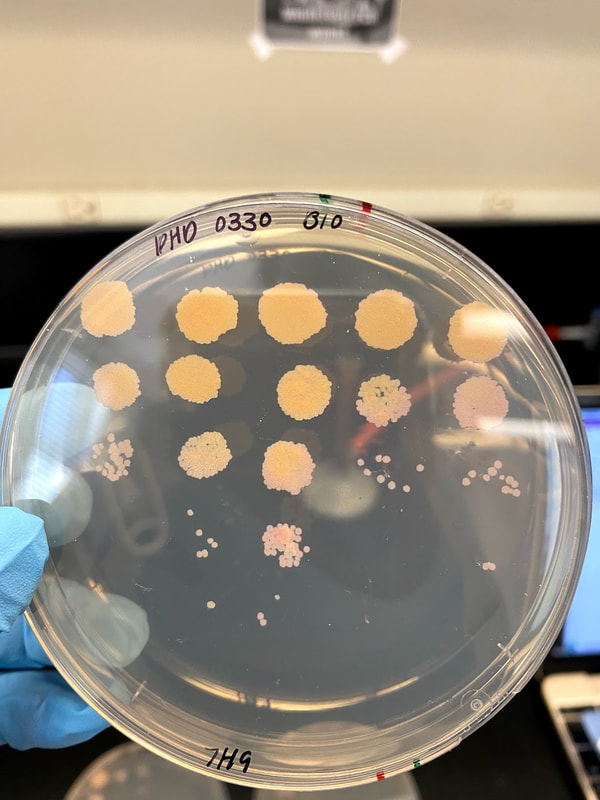
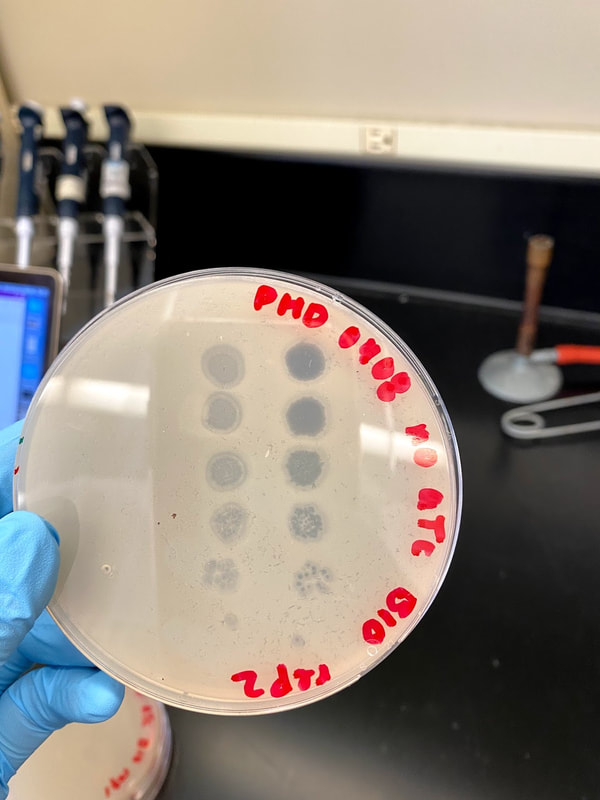
Through Lehigh's SEA PHAGES program, I have isolated and published my own novel bacteriophage, named Tiri.
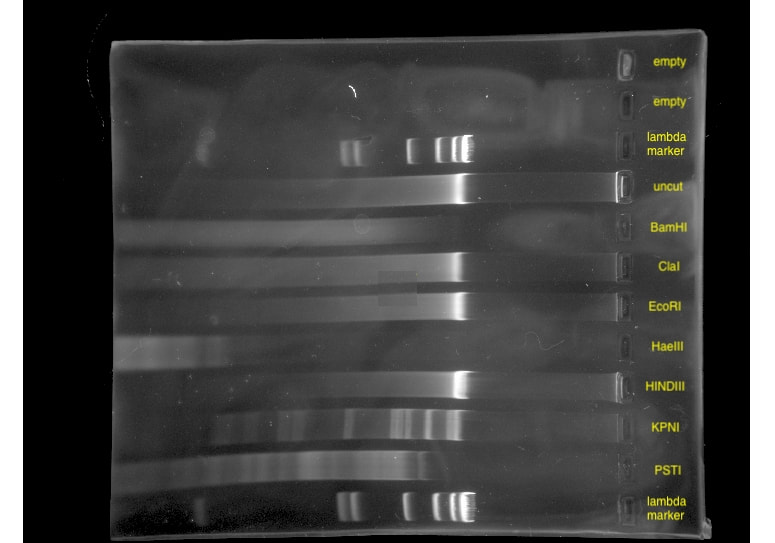
Techniques that I employed to isolate this phage included gel electrophoresis, plaque assays, serial dilutions, aseptic technique, and plating and spotting of bacterial cultures. Phage Tiri was sequenced in 2022 and determined to be a K1 phage, as per my prediction!
Techniques that I employed to isolate this phage included gel electrophoresis, plaque assays, serial dilutions, aseptic technique, and plating and spotting of bacterial cultures. Phage Tiri was sequenced in 2022 and determined to be a K1 phage, as per my prediction!
I have also used bioinformatics programs (DNA Master, phamerator, BLAST, HHpred) to analyze genomes and determine gene functions of multiple bacteriophage sequences. These results are currently in review for publication.
In spring 2021, through the SEA GENES program, I worked on amplifying and characterizing multiple genes of Mycobacteriophage Butters. I was able to successfully amplify and create primers for two genes that had never been amplified before, although they were worked on by various students for three years prior. For this work, I ran PCR reactions, inoculated bacterial cultures, electroporated and transformed competent cells, and conducted phenotypic assays.
I also conducted independent research on the mechanisms of bacteriophage genome recombination. This project utilizes the technique of BRED to introduce mutations into bacteriophage genomes in an attempt to elucidate the factors (genes) that contribute to the creation of recombinant phages. This research has implications in the safety and viability of phages as alternatives to antibiotics in the case of superbugs.
This work was presented as a talk at the SEA (Science Education Alliance) Symposium in April 2021, and the following abstracts were also presented:
- Mohammed H.T., J. Korenberg, A. Edmundson, B. Bachiashvili, P. Demetriades, R. Schmidt, C. Mageeney, M. Kenna, V.C. Ware. (2021). Drivers of Recombination between Phage Genomes: Mycobacteriophage Butters prophage-mediated constraints on Island3 infection of Mycobacterium smegmatis. (Abstract)
- Andryshak S., A. Boyer, G. Ciabbattoni, H. Cho, N. Clarke, P. Demetriades, N. DesGranges, B. Dubyna, M. Dunterman, E. El-Halawani, M. Fehrenbach, G. Forsyth, A. Fucile, S. Haab, A. Iftikhar, R. Kyle, R. Milelli, C. Murphy, N. Novak, R. O’Toole, R. Roberts, S. Rosenthal, A. Sanchez, J. Skrapits, M. Sorbello, G. Stephenson, N. Weaver, and M. Kuchka. (2021) Elucidating Mycobacteriophage Butters Gene Functions with the SEA-GENES Research Project at Lehigh. (Abstract)
- Bachiashvili B., C. Cosgrove, P. Demetriades, L. Duffany, A. Edmundson, I. Foster, A. Fucile, E. Greenholt, S. Haab, R. Milelli, A. Salcedo, J. Shah, G. Stephenson, K. Sun, N. Yellamelli, T. Zhang, H. Mohammed, M. Kenna, V. Ware. (2021). Bioinformatics Discoveries: Investigating Mycobacteriophage Genomic Relationships and Immunity Mechanisms. (Abstract)
In the spring of 2022, I worked on isolating a second novel phage, which I took from marsh soil at Island Beach State Park (near my mud snails)!
At the SEA Symposium in April 2022, the following abstracts featuring my work were presented:
- Anandani R., B. Bachiashvili, P. Demetriades, A. Fucile, E. Greenholt, S. Haab, Q. Klessel, R. Mansi, R. Milelli, J. Schalet, J. Shah, J. Siani, T. Zhang, S. Mensah, H. Mohammed, M. Kenna, M. Kuchka, V. Ware. (2022). Deep SEA Diving: An Advanced Phage Genetics Course. (Abstract)
- Anandani R., R. Antar, K. Dayi, P. Demetriades, F. Demissie, S. Haggert, Q. Klessel, K. Parsa, J. Schalet, L. Shebby, J. Siani, W. Yaeger, N. Zvenyika, H. Mohammed, R. Schmidt, M. Kenna, V. Ware. (2022). The Power of DOGEMS: Analysis of Several Mycobacteriophage Genomes. (Abstract)